MCAT Biochemistry Review Summary
Gold Standard MCAT Biochemistry Review Summary ('cheat sheet' notes)
This MCAT Biochemistry Review Summary Page will draw your attention to the most important and helpful points and helps to understand vocabulary, definitions, and relationships in biochemistry. In some respects, learning this "language" of biochemistry will decrease your need to memorize information and increase your understanding of MCAT Biochemistry content. Unlike most undergraduate Biochemistry exams, MCAT Biochemistry is far more likely to ask reasoning questions than to ask for the intermediate or enzyme in a pathway that you are expected to review.
We have placed a couple of practice questions on this page, and you can access more practice questions in our free MCAT practice test or our many full-length MCAT practice tests. You may also be interested in Gold Standard's free premium app: MCAT Biochemistry for iPhone, MCAT Biochemistry for Android.
For a detailed list of MCAT Biochemistry topics see our MCAT topics list.
Download the Gold Standard MCAT Biochemistry Review Summary PDF or scroll down and click on an image below to view the summaries.
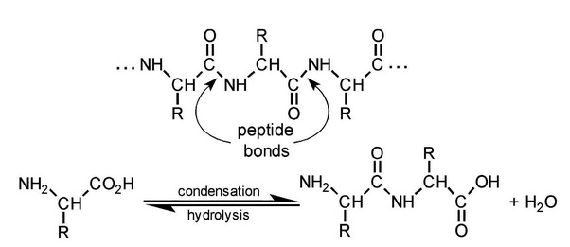
MCAT Biochemistry - Macromolecules, Kinetics, Thermodynamics, & The Plasma Membrane
| Building Block | Polymerizes to form . . . | Chemical bonds | Macromolecule |
| Monomers | Dimer, trimer, tetramer, oligomers, etc. | Covalent bonds* | Polymer |
| Amino acids | Dipeptide, tripeptide, tetra/oligopeptide, etc. | Peptide bonds | Polypeptide, protein |
| Monosaccharides ('simple sugars'**) | Disaccharide, tri/tetra/oligosaccharide, etc. | Glycosidic bonds | Polysaccharide |
| Nucleotides | Nucleotide dimer, tri/tetra/oligomer, etc. | Phosphodiester bonds | Polynucleotides, nucleic acids |
*There are exceptions. For example, in certain circumstances polypeptides are considered monomers and they may bond non-covalently to form dimers (i.e. higher orders of protein structure).
**Note that disaccharides are also sugars (i.e. sucrose is a glucose-fructose dimer known as 'table sugar'; lactose is a glucose-galactose known as 'milk sugar').
Macromolecules: Summary I - Amino Acids and Proteins
-
The current MCAT regularly has questions which require previous knowledge of the structures, features (including changes in charge with pH), 3- and 1-letter abbreviations of the 20 common protein-generating amino acids, etc. You can consider the following your MCAT Biochemistry cheat sheet (i.e. memory aid for quick reference, but obviously not to be used surreptitiously).
Note: Unless mentioned otherwise, the following images are excerpts from the Gold Standard MCAT Biochemistry ebook.
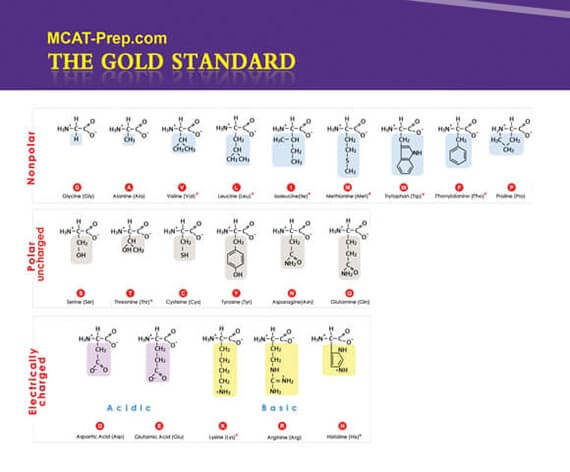
Ala A Alanine Arg R Arginine Asn N Asparagine Asp D Aspartic acid Cys C Cysteine Gln Q Glutamine Glu E Glutamic acid Gly G Glycine His H Histidine Ile I Isoleucine Leu L Leucine Lys K Lysine Met M Methionine Phe F Phenylalanine Pro P Proline Ser S Serine Thr T Threonine Trp W Tryptophan Tyr Y Tyrosine Val V Valine Ready to test your amino acid and protein knowledge (and more)?
Sign up for our FREE full-length mock exam and challenge yourself on key MCAT topics. Get instant scores, detailed explanations, and access to helpful science videos. Sign up here: Free MCAT practice test
Helpful Mnemonics for One-Letter Amino Acid Abbreviations on the MCAT
-
Alanine, aRginine, asparagiNe, asparDic acid [aspartic acid], Cysteine, Qutamine [glutamine], glutamEc acid [glutamic acid], Glycine, Histidine, Isoleucine, Leucine, Kysine [lysine], Methionine, Fenylalanine [phenylalanine], Proline, Serine, Threonine, tWyptophan [tryptophan], tYrosine, Valine
Nonpolar Amino Acids = G A P V W L I M F or Gap V.W. Lymph
(lymph is important in fatty acid transport . . . think fatty acid tails are nonpolar)
Polar Uncharged Amino Acids = S T Y C N Q or Stick Nick
(stick like a "pole" . . . think polar)
Electric Amino Acids = D E H K R or Dee Hicker
(dee hicker like deelectric . . . think electric)
To be more precise, for the Polar Charged Amino Acids:
Dee Negative, Hicker Positive, D(-) E(-) H(+) K(+) R(+)
General Principles of Proteins
-
- Primary structure
- Sequence of amino acids encoded by DNA and linked to each other through covalent bonds (including disulfide bonds)
- Secondary structure
- Orderly inter- or intramolecular hydrogen bonding of the protein chain
- Usually organized in a stable ɑ-helix or a β-pleated sheet
- Tertiary structure
- Further folding of the protein molecule onto itself
- 3D shape (spatial organization) of an entire protein molecule
- Quaternary structure
- 2 or more protein chains bonded together by noncovalent bonds
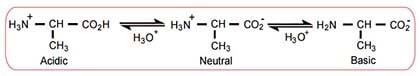
- Isoelectric point (pI)
- pH at which a given amino acid will be neutral (have no net charge)
- Average of the 2 pKa values of an amino acid (depending on the dissociated group)
- Both amino acids and proteins are least soluble at their isoelectric points
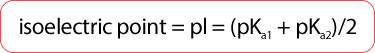
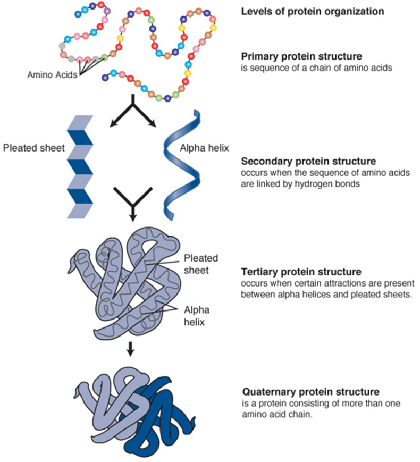
Levels of Protein Organization
[Wikipedia, Creative Commons]
- Primary structure
Macromolecules: Summary II - Carbohydrates
-
- Compounds consisting of aldehydes and ketones with general formula Cm(H2O)n
- Monosaccharides = simplest carbohydrate unit
- Can be named according to number of carbons it contains
- Common names: triose (3 carbons), tetrose (4 carbons), pentose (5 carbons), hexose (6 carbons)
- Note: '-ose' suffix denotes sugar
- Aldose = aldehyde sugar
- Ketose = ketone sugar
- Disaccharides = two sugars bound together
- Polysaccharides = long sugar chains for glucose storage (i.e. glycogen in animals, starch in plants)
- Absolute configuration
- Describes spatial arrangement of atoms or groups around a chiral molecule
- Assign priority to substituents of the chiral carbon
- The higher the atomic number, the higher the priority
- Orient the molecule with the lowest priority substituent in the back
- If the priorities increase in a clockwise direction = R
- If the priorities decrease in a clockwise direction = S
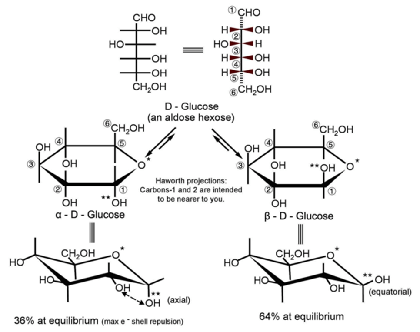
- Cyclic structure
- Monosaccharides can undergo intramolecular reactions to form ring structures
- Cyclic sugars are stable in solution and can form two types of rings:
- Six-membered rings (6 carbons in ring) = pyranose
- Five-membered rings (5 carbons in ring) = furanose
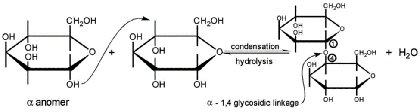
- Glycosidic linkages
- Covalent bonds between monosaccharides and alcohols
- When alcohol is another monosaccharide, produces a disaccharide
- i.e. sucrose, lactose, maltose, cellobiose
- Linkage between C1 on the first sugar and C2 on the second sugar = 1,2 linkage
- Linkage between C1 on the first sugar and C4 on the second sugar = 1,4 linkage
- Linkage between C1 on the first sugar and C6 on the second sugar = 1,6 linkage
- May also be classified as alpha (α) or beta (β):
- Hydroxyl group on C1 oriented up = beta (βirds fly in the sky)
- Hydroxyl group on C1 oriented down = alpha (α - fish - swim in the sea)
- Epimers vs. Anomers
- Epimers - differ in absolute configuration at only one carbon (R or S)
- Anomers - differ in configuration at the hemiacetal/acetal (= anomeric) carbon
- Note: Conversion between anomers = mutarotation
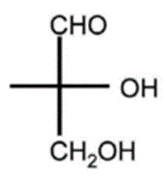
D, R-glyceraldehyde
Macromolecules: Summary III - Fatty Acids
-
- Amphipathic molecules possessing a polar carboxylate-head group and nonpolar hydrocarbon tail
- Saturated fatty acids
- Maximum hydrogens - stack together
- No double bonds
- Solids at room temperature (= fats; example: butter)
- Higher melting point, boiling point, and Van der Waals
- Unsaturated fatty acids
- 1 or more double bonds - bent or kinked structure
- Liquids at room temperature (= oils; example: olive oil)
- Exist in body in 3 common forms:
- Triglycerides (also known as triacylglycerols)
- 3 fatty acids esterified to glycerol
- Nonpolar tails - used for energy storage
- Digested by lipase
- With low food intake, broken down by liver to form ketone bodies
- Phospholipids
- Fatty acids bound to glycerol, phosphate, and other groups
- Amphipathic (hydrophilic + hydrophobic components) - used for cell membranes
- Cholesterol
- Fatty acids in ring form [isoprene units derived from acetyl-CoA cyclize to form a four-ringed structure]
- Cholesterol is amphipathic. The ring is nonpolar and the OH-head group is polar.
- Used for steroid hormones, bile, membrane fluidity
- Triglycerides (also known as triacylglycerols)
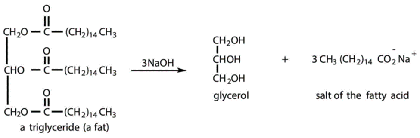
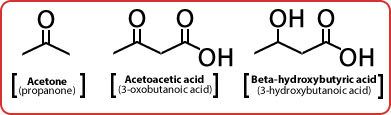
The Three Ketone Bodies
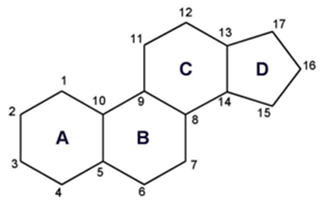
Basic Ring Structure of Steroids
Macromolecules: Summary IV - Nucleic Acids
-
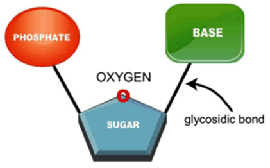
- Nucleotides (also known as nucleoside phosphates)
- are composed of a 5-carbon sugar, a nitrogenous base, and an inorganic phosphate (a glycosidic bond links the nitrogenous base to the sugar)
- are attached in sequence via phosphodiester bonds to form nucleic acids
- form the sugar phosphate (negatively-charged) backbone of nucleic acids
- Nucleosides
- are composed of a nitrogenous base and a 5-carbon sugar
- are attached to an inorganic phosphate to form nucleotides
- Nitrogenous Bases
- Purines
- 2 rings - adenine (A) and guanine (G) - mnemonic: Pure As Gold
- Pyrimidines
- 1 ring - cytosine (C), uracil (U), thymine (T) - mnemonic: CUT the Py
- Purines
Note the direction of hydrogen bonding in the diagram below:
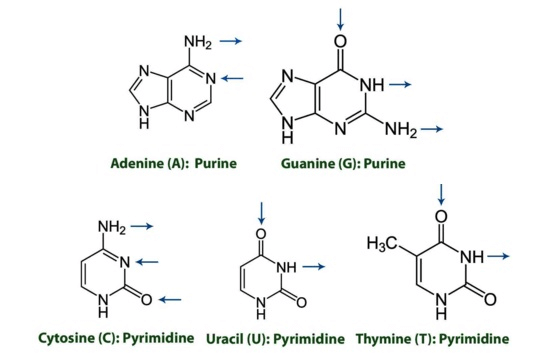
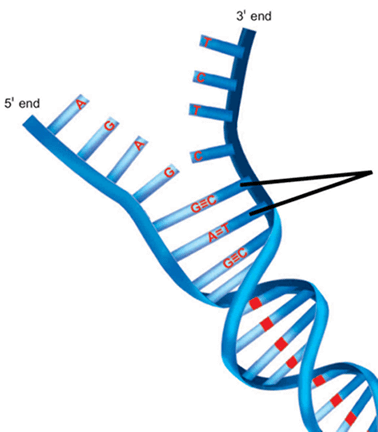
- Watson–Crick model of DNA
- Double helical structure composed of 2 complementary and anti-parallel DNA strands held together by hydrogen bonds between nitrogenous base pairs
- A binds T with 2 hydrogen bonds; C binds G with 3 hydrogen bonds
- Mnemonic: All Tigers and Cats Growl
- Nitrogenous bases project to center of double helix to hydrogen bond with each other (think of double helix as a winding staircase - each stair represents a nitrogenous base pair keeping the helix shape intact)
- Double helical structure composed of 2 complementary and anti-parallel DNA strands held together by hydrogen bonds between nitrogenous base pairs
Deoxyribonucleic Acid (DNA) Ribonucleic Acid (RNA) Contains the genetic information of the cell - transfers that genetic information Helps DNA transfer genetic information for creation of proteins Made from deoxyribose Made from ribose Double-stranded Mostly single-stranded Adenine binds Thymine
Cytosine binds GuanineAdenine binds Uracil
Cytosine binds GuanineFound in nucleus and mitochondria Found in nucleus, cytoplasm, and ribosomes Replicated by DNA polymerases Transcribed from DNA - DNA Denaturation
- Separation of DNA strands due to high temperatures, chemicals (i.e. urea), or UV radiation
- Occurs when hydrogen bonds between base pairs are broken
- Separated strands are still complementary
- DNA Reannealing (also known as hybridization)
- Single, complementary strands stick tightly and reform double-stranded DNA
- Single strands of DNA can only hybridize with each other if they are complementary
- Nucleotides (also known as nucleoside phosphates)
Macromolecules: Summary V - Metabolism & Nomenclature
-
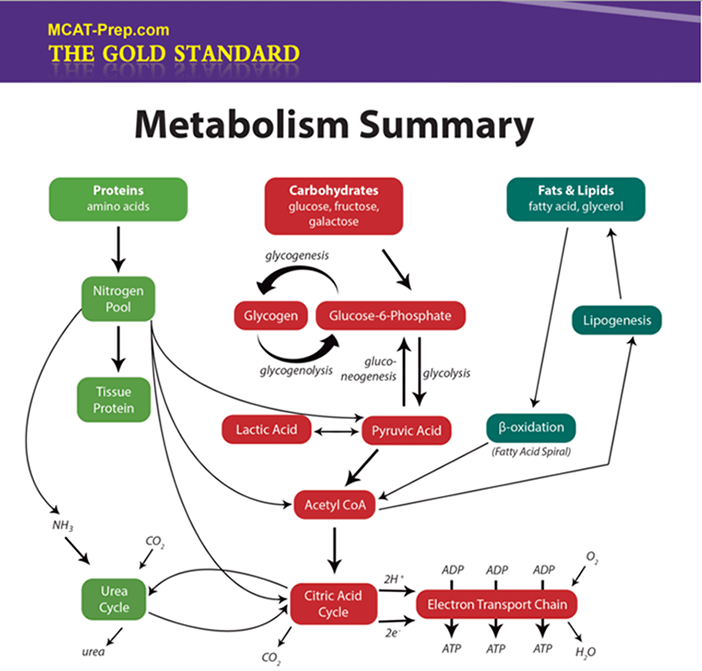

MCAT Biochemistry: Kinetics Summary I
-
- Kinetics tell how fast a reaction will occur
- Catalysts
- speed up reactions by speeding up the rate-determining step or providing a different route to the products; lower the energy of activation
- remain unchanged at the end of the reaction
- are not included in the overall reaction equation
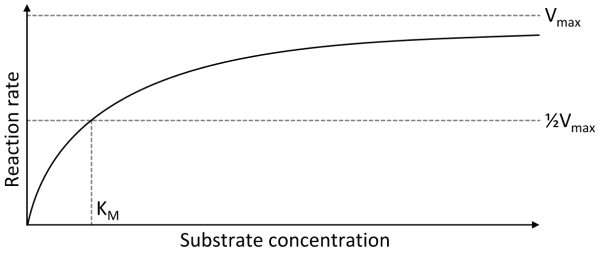
- Michaelis-Menten
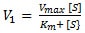
- V0 = initial velocity (reaction rate)
- Vmax = maximum reaction rate
- [S] = concentration of substrate
- Km = amount of substrate needed for enzyme to obtain half of its maximum rate of reaction (Michaelis-Menten constant)
- Low Km = high affinity for substrate
- High Km = low affinity for substrate
- Ka = association constant of the enzyme-substrate complex
- Low Ka = low affinity for substrate (enzyme-substrate complex is less stable)
- High Ka = high affinity for substrate (enzyme-substrate complex is more stable)
- Kd = dissociation constant of the enzyme-substrate complex
- Low Kd = high affinity for substrate (enzyme-substrate complex is more stable)
- High Kd = low affinity for substrate (enzyme-substrate complex is less stable)
- kcat = number of substrate molecules each enzyme converts to product per unit time (turnover number)
- Low kcat = enzyme-substrate complex converts less of substrate it binds into product
- High kcat = enzyme-substrate complex converts more of substrate it binds into product
- Feedback Regulation
- Positive Feedback - promotes or enhances reactions
- Negative Feedback - inhibits or reduces reactions
- Cooperativity (also known as positive or negative cooperative binding)
- Occurs when binding of Substrate B to Substrate A increases (positive) or decreases (negative) activity of Substrate A
- Michaelis-Menten kinetics do not apply to enzymes engaging in cooperative binding
MCAT Biochemistry: Kinetics Summary II - Enzymes
-
- Competitive Inhibitors
- Compete with the substrate in binding to the enzyme's active site
- Are structurally similar to the substrate
- Bind to free enzyme (= E) only forming an unreactive enzyme-inhibitor complex (= EI)
- Cannot bind to the enzyme at the same time as the substrate
- Can only bind if the substrate has not bound
- Are reversible with increasing [S]
- Increase Km and have no effect on Vmax
- Uncompetitive Inhibitors
- Bind to the enzyme-substrate complex (= ES) only forming an unreactive enzyme-inhibitor-substrate complex (= EIS)
- Can only bind if the substrate has bound
- Are reversible with decreasing [S]
- Decrease Km and decrease Vmax
- Mixed Inhibitors
- Bind to the enzyme at a site other than the active site
- Bind to free enzyme (= E) or the enzyme-substrate complex (= ES) forming an unreactive enzyme-inhibitor complex (= EI) or enzyme-inhibitor-substrate complex (= EIS), respectively
- Can bind whether or not the substrate has bound
- Produce a conformational change in the enzyme - binding of the inhibitor reduces the substrate's affinity for the active site
- Are reduced, but not reversible with increasing [S]
- Increase or decrease Km and decrease Vmax
- Noncompetitive Inhibitors
- Do not compete with the substrate in binding to the enzyme's active site
- Bind to the enzyme at a site other than the active site
- Bind to free enzyme (= E) or the enzyme-substrate complex (= ES) forming an unreactive enzyme-inhibitor complex (= EI) or enzyme-inhibitor-substrate complex (= EIS), respectively
- Can bind whether or not the substrate has bound
- Are not reduced or reversible with increasing [S] - must remove inhibitor
- Decrease Vmax and have no effect on Km
Note: Noncompetitive inhibition is sometimes considered a special case of mixed inhibition. Noncompetitive inhibitors have the same affinity for free enzyme (= E) and the enzyme-substrate complex (= ES) whereas mixed inhibitors tend to have a higher affinity for either free enzyme (= E) or the enzyme-substrate complex (= ES).
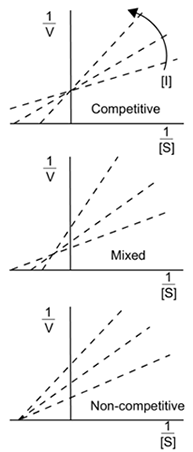
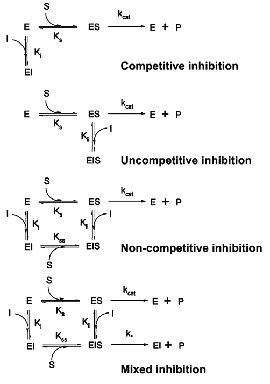
[Wikipedia, Creative Commons]
Inhibitor Binds Reversible? Effect Competitive Free enzyme (E) Yes with ↑[S] Increases Km Uncompetitive Enzyme-substrate complex (ES) Yes with ↓[S] Decreases Km and Vmax Mixed Free enzyme (E) or enzyme-substrate complex (ES) Reduced with ↑[S] Increases or decreases Km and decreases Vmax Noncompetitive Free enzyme (E) or enzyme-substrate complex (ES) Yes with removal of inhibitor Decreases Vmax KoMpetitive INhibition = KM INcrease (Vmax is unchanged)
NOn-KoMpetitive INhibition = NO KM INcrease (but Vmax is decrease)
Uncompetitive Inhibition = BOTH Km and Vmax decrease
Mnemonic for Remembering the Effects of Enzyme Inhibitors
- Ki = binding affinity of the inhibitor
- Ki > 1 - inhibitor has higher affinity for enzyme than substrate
- Ki < 1 - inhibitor has lower affinity for enzyme than substrate
- IC50 = half-maximal inhibitory concentration
- Tells how much of a drug is needed to inhibit a biological process by 50%
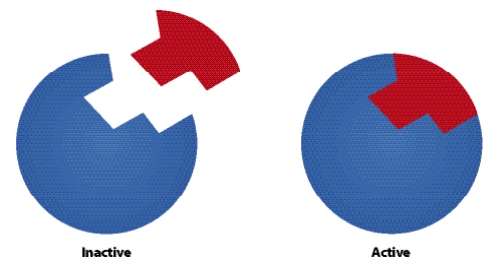
- Allosteric enzymes have 2 configurations:
- Active state - catalyzes reactions
- Inactive state - cannot catalyze reactions
- Zymogens are inactive enzymes that must be cleaved to become active
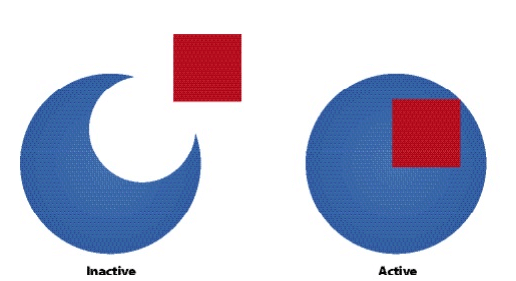
- Competitive Inhibitors
MCAT Biochemistry: Thermodynamics Summary
-
- Thermodynamics tell if a reaction will occur
- Gibbs Free Energy (ΔG) - tells nothing about rate of reaction, only its spontaneity
- ΔG = ΔH - TΔS
- Negative ΔG = exergonic = spontaneous
- ΔG = 0 = equilibrium
- Positive ΔG = endergonic = nonspontaneous
- Equilibrium constant (Keq)
- Keq = products/reactants
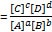 for a reaction A + B → C + D
for a reaction A + B → C + D
- Note: a, b, c, and d are stoichiometries of the substrates and products, respectively
- Keq > 1 - reaction favors products
- Keq < 1 - reaction favors reactants
- Keq = products/reactants
- Gibbs Free Energy at standard state (ΔGo) = 298K (25oC), 1 atm, pure solids/liquids, 1M solutions
- ΔGo < 0 and Keq > 1 . . . products are favored
- ΔGo = 0 and Keq = 1 . . . products are approx. equal to reactants
- ΔGo > 0 and Keq < 1 . . . reactants are favored
- Le Chatelier's Principle
- Reaction in equilibrium will change direction according to applied stress
- Stress could be a change in concentration, pressure, temperature, or volume
- Enthalpy
- Measure of heat energy absorbed or released when bonds are broken or formed, respectively, during a reaction run at constant pressure
- Negative ΔH - exothermic - bond(s) formed - energy released
- Positive ΔH - endothermic - bond(s) broken - energy absorbed
ΔH ΔS ...ΔG Reaction Spontaneity - + - Spontaneous at all temperatures - - - or + Spontaneous at low temperatures where ΔH outweighs TΔS
Nonspontaneous at high temperatures where TΔS outweighs ΔH
+ - + Nonspontaneous at all temperatures + + - or + Spontaneous at high temperatures where TΔS outweighs ΔH
Nonspontaneous at low temperatures where ΔH outweighs TΔS
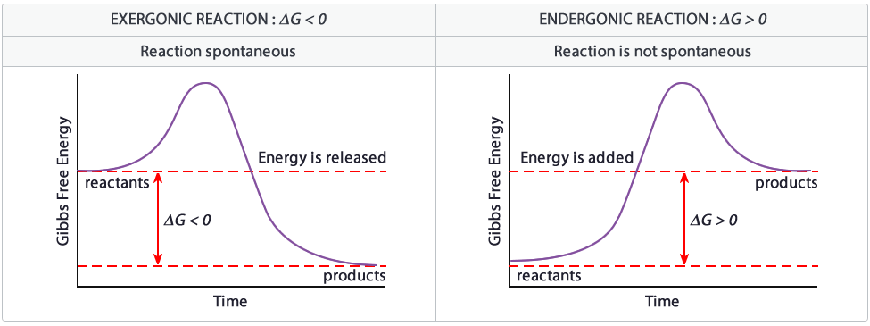
Plasma Membrane: Summary I
-
- A phospholipid bilayer that encloses cells
- Outer region - hydrophilic, polar, phosphoric acids (= heads)
- Inner region - hydrophobic, nonpolar, fatty acids (= tails)
- Cholesterol found within membrane for structural support and as a fluidity buffer
- Transmembrane proteins assist in solute transport across membranes
Process (solute type) Concentration Gradient Requires Protein? Requires Energy? Diffusion (small nonpolar) High to Low No No – passive Osmosis (water only) High to Low No No – passive
Facilitated Transport (large nonpolar) High to Low Yes No – passive Active Transport (polar/ions) Low to High Yes Yes – active (ATP)
Note: Difference in polarity between (+) outside and (-) inside along with solute transport creates a membrane potential across the cell membrane.
Osmotic pressure = pressure required to resist movement of water through a semipermeable membrane due to concentration gradient
Plasma Membrane: Summary II
-
- Exocytosis
- Vesicle inside cell fuses with cell membrane and releases contents to outside
- Endocytosis
- Cell membrane bends inward to form vesicle around particles or liquids
- If large particles = phagocytosis
- If small particles or liquids = pinocytosis
- Cell membrane bends inward to form vesicle around particles or liquids
- Note: Contents never cross cell membrane in either process
- Biosignaling
- Ion Channels - allow for passive or active transport of ions; two types:
- Voltage-gated - transmembrane proteins activated by changes in membrane potential
- Ligand-gated - transmembrane proteins activated by binding of a specific ligand
- Ion Channels - allow for passive or active transport of ions; two types:
- Receptor enzymes (also known as enzyme-linked receptors)
- Transmembrane proteins
- Binding of ligand on extracellular side produces activity on the intracellular side
- G protein-coupled receptors
- Integral membrane proteins
- Sensing of molecules outside cell activates response inside cell
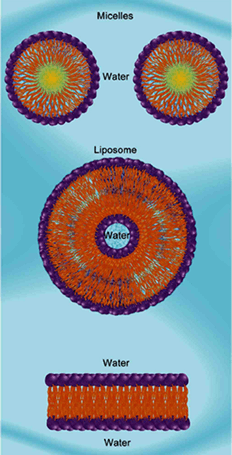
Amphipathic molecules arranged in micelles, a liposome and a bilipid layer [= plasma membrane]
- Exocytosis
Biotechnology: Summary I
-
- DNA recombination
- Involves combining DNA segments or genes from different sources
- Foreign DNA can come from another DNA molecule, a chromosome, or a complete organism
- Uses restriction enzymes from bacteria that cut pieces of DNA at specific recognition sites (= nucleotide sequences) along the DNA strand
- When a double-stranded DNA segment is cut, sticky ends are produced
- Sticky ends = unpaired, single-strand extensions of DNA that are ready to bind with complementary codons (= sequence of 3 adjacent nucleotides)
- The cut pieces of DNA = restriction fragments
- Restriction fragments are then inserted into plasmids
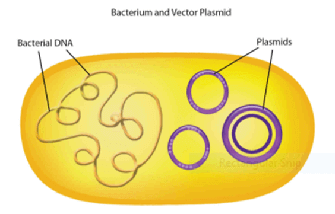
- Plasmids (also known as replicons)
- Circular pieces of DNA that replicate independently of chromosomal DNA
- Cut with the same restriction enzymes as the restriction fragment to generate the same sticky ends
- Plasmids (also known as replicons)
- Plasmids are introduced into the bacteria via transformation
- Restriction fragments and linearized plasmids are joined together by DNA ligase to form a recombinant plasmid
- Special application
- DNA recombination makes it possible for bacteria to make many copies of recombinant plasmids and produce proteins that are not native to its species (e.g. bacteria with recombinant DNA producing insulin to treat diabetes)
- More on restriction enzymes (also known as restriction endonucleases - endo = internal; nuclease = cut DNA)
- Naturally found in bacteria and archaea
- Serve as the organism's own natural defense mechanism
- Enzymes cleave foreign DNA at specific recognition sites when it enters a bacterium
- The organism's own DNA is protected by selective methylation at these sites via methylase enzymes
- Come in 3 types - classified by their structure and how they cleave their DNA substrate
- Type I, II, and III – Type II is the most common
- Type II restriction enzymes
- Cleave within or near specific recognition sites (short, specific, palindromic DNA sequences where cleavage occurs under the right conditions)
- Note: palindromic sequences read the same forward or backward (e.g. RACE CAR)
- Gene cloning
- Involves the use of a cloning vector (e.g. plasmid, bacteriophage, virus, etc.)
- All cloning vectors must contain at least one unique cloning site
- That cloning site must be cleavable by a restriction enzyme as well as a place where a gene of interest can be inserted
- Multiple Cloning Site
- An artificially-engineered region of several cloning sites lumped together
- Found in many commercial and artificially modified cloning vectors
- Allows the researcher to pick restriction sites that:
- are unique to the vector
- ensure that translocation occurs in-frame
- Origin of replication (ori) = allows cloning vectors to self-replicate
- Special applications
- Cloning vectors are often modified to carry genes that provide resistance to antibiotics, like ampicillin or kanamycin, which allows vectors to be used for selecting certain bacteria
- Bacteriophage vectors have high transfection (= introduction of nucleic acids into cells) rates and are easy to screen which makes them useful for large-scale applications (e.g. genomic DNA screening)
- DNA libraries
- are collections of recombinant DNA
- can consist of (1) genomic DNA or (2) cDNA
- Genomic DNA library
- Contains recombinant DNA that represents the entire genome of an organism
- How it is generated:
- Partially digest genomic DNA with restriction enzymes to ensure representation of all genes
- Digests bacteriophage vector with the same enzyme
- Ligate digested genomic DNA with vector - pack both into bacteriophages
- Infect densely populated E.coli with bacteriophages
- Isolate cleared areas of lysed cells (= plaques)
- cDNA (= complementary DNA) library
- Contains double-stranded (= dsDNA) that is reverse transcribed from messenger RNA (= mRNA)
- How it is generated (in eukaryotes):
- Purify mRNA by applying it to an oligodeoxythymidylate (oligo dT) cellulose filter which binds the mRNA's poly A-tail
- Synthesize cDNA using oligo dT as a primer
- Cut mRNA with RNA-nicking RNase H and DNA polymerase I uses the fragments as primers to synthesize cDNA
- Digest resulting double-stranded cDNA and insert it into vectors
- Create phage library as discussed for genomic DNA library
- Unlike genomic DNA libraries, an advantage of cDNA libraries is that they do not include introns or flanking sequences
- Locating a gene of interest
- Use a hybridization probe
- A molecule that recognizes a specific DNA sequence or protein product labeled with a compound that allows for easy detection (e.g. radioactive isotope, reactive enzyme that produces color, or fluorescent dye)
- Hybridize the probe and the DNA library
- The sensitivity or stringency of the hybridization reaction can be increased or decreased by changing experimental variables (e.g. salt concentration, temperature)
- Isolate detected positive hits from phage plates
- Identify and isolate the gene of interest
- Amplify the gene using polymerase chain reaction (PCR)
- Use a hybridization probe
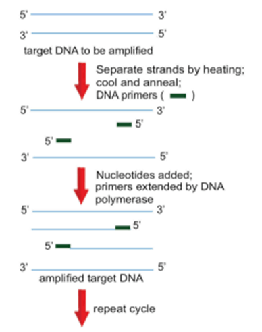
- Polymerase Chain Reaction (PCR)
- Allows for the rapid amplification of any DNA fragment without purification
- DNA primers flank the specific sequence to be amplified
- Primers are then extended to the end of the DNA molecule with the use of a heat-resistant DNA polymerase
- Newly synthesized DNA strand is then used as the template to undergo another round of replication
- Technique:
- Melt target DNA into two single strands by heating the reaction mixture to approximately 94 °C
- Cool the mixture rapidly to anneal the primers to their specific locations
- Note: anneal = the sticking together of complementary single strands via the formation of hydrogen bonding between base pairs
- Raise the temperature to 72°C to allow DNA polymerase to add nucleotides until the entire complementary strand of the template is completed
- Repeat the cycle to make more copies
- Special applications:
- Sex determination
- Requires amplification of intron 1 of amelogenin gene
- Amelogenin gene, found on X-Y homologous chromosomes, contains a 184 base pair deletion on the Y homologue
- That deletion allows females to be distinguished from males in that males will have two different sizes of amplified DNA whereas females will have one unique size
- Introduce specific mutations into genes for research purposes such as:
- Deleting entire fragments
- Changing specific amino acids
- Adding peptide tags (= peptide sequences tagged onto a protein)
- Sex determination
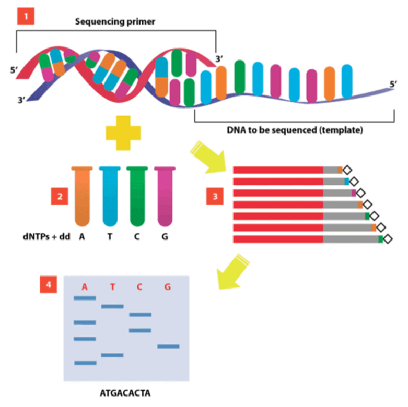
- DNA Sequencing
- Method by which to ensure that the gene insert of interest does not contain any mistakes
- Sanger method of DNA sequencing
- Most commonly used sequencing technique
- Makes four preparations of DNA
- Each preparation is incubated with a labeled sequencing primer, DNA polymerase, all 4 DNA triphosphates, and 1 of the nucleotides in its dideoxy form (e.g. ddATP, ddCTP, ddGTP, or ddTTP)
- Technique:
- New strands of DNA are synthesized using PCR beginning at the primer and continuing on until a dideoxynucleotide (ddNTP) is inserted at random
- This results in a mixture of DNA fragments with varying lengths that must be separated by gel electrophoresis and read one nucleotide at a time
- DNA recombination
Biotechnology: Summary II
-
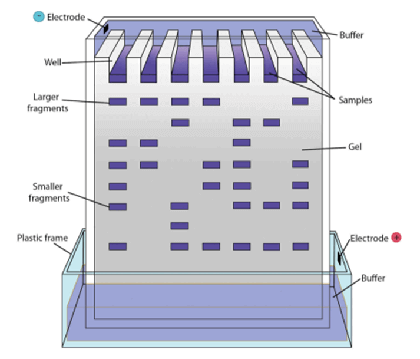
- Gel electrophoresis
- Method of separating DNA fragments of differing lengths based on their size
- Technique:
- DNA fragments are passed through an electrically-charged agarose gel
- Since DNA is negatively charged, it moves towards the anode (= positive electrode)
- Shorter fragments move faster than the longer fragments and can be visualized as a banding pattern using autoradiography techniques
- Southern blotting
- Method of transferring DNA fragments from the electrophoresis agarose gel onto filter paper where they are identified with probes
- Technique:
- DNA is digested in a mixture with restriction endonucleases to cut out specific pieces of DNA
- The DNA fragments are then subjected to gel electrophoresis
- Separated fragments are bathed in an alkaline solution where they immediately begin to denature
- Fragments are then blotted onto nitrocellulose paper and incubated with a specific probe whose location can be visualized with autoradiography
- Northern blotting
- Method of detecting specific sequences of RNA by hybridization with cDNA
- Western blotting
- Method used to identify specific amino acid sequences in proteins
- Proteins are separated by gel electrophoresis on an SDS-PAGE gel and detected using labeled antibodies specific to the protein or part of the protein

Mnemonic for Remembering the Blotting Techniques
- DNA microarrays (= DNA chip or biochip or "laboratory-on-a-chip")
- Determine which genes are active or inactive in different cell types
- Evolved from Southern blotting
- Can be used to genotype multiple regions of a genome
- Created by robotic machines that arrange small amounts of hundreds of thousands of gene sequences on a single microscope slide
- Sequences can be a short section of a gene or other DNA element that is used to hybridize a cDNA or cRNA (also called antisense RNA) sample
- Hybridization is usually observed and quantified by the detection of a fluorescent tag
- Protein Quantification
- Quantitative analysis techniques allow researchers to determine basic information about the expressed product (e.g. concentration, amount, size)
- SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis)
- Method of separating proteins according to their electrophoretic mobility
- SDS is an anionic (= negatively charged) detergent with the following effects: (1) denaturing proteins and (2) giving an additional negative charge to the now-linearized proteins
- In most proteins, the binding of SDS to the polypeptide chain gives an even distribution of charge per unit mass, thus fractionation will approximate size during electrophoresis
- Column Chromatography
- Method of purifying proteins according to their size, charge, hydrophobicity, and more
- Affinity Chromatography
- Allows the rapid purification of biological molecules based on certain affinities
- Examples: affinities between antigen and antibody, enzyme and substrate, or genetically engineered protein tags and specific matrices
- Ion Chromatography
- Allows the purification of biological molecules based on overall charge:
- Cation Chromatography - selectively binds positively-charged molecules
- Anion Chromatography - selectively binds negatively-charged molecules
- Proteins are removed by washing (= eluting) using increasing concentrations of salt
- Allows the purification of biological molecules based on overall charge:
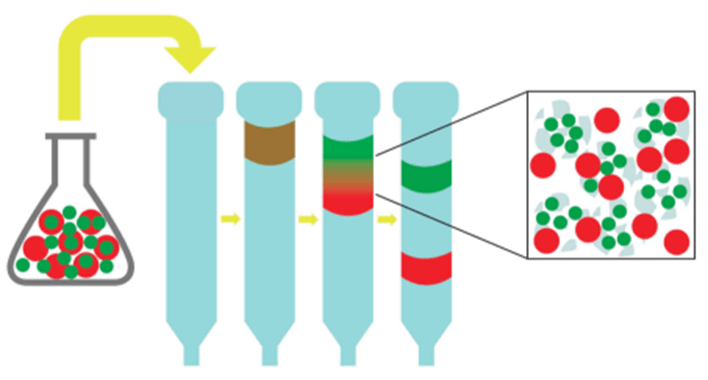
- Size-Exclusion Chromatography (SEC) (also known as gel filtration)
- Allows the purification of biological molecules based on size
- A mixed protein solution is applied to the top of a long column packed with porous polymer beads
- As the solution filters through the beads, smaller molecules are slowed down by the pores in the beads while larger molecules are allowed to flowed through at a faster rate - thus, larger molecules are eluted first
MCAT Biochemistry Practice Questions
-
These MCAT Biochemistry practice questions are taken from our Gold Standard MCAT Question Bank that contains over 4000 practice questions and growing. These questions were chosen to give you a sense as to the reasoning and knowledge required in Biochemistry for the current MCAT exam. It would be better to complete some part of your review before attempting these practice questions. Good luck!
MCAT Biochemistry Practice Question 1
In order to become active, a Map kinase has to undergo phosphorylation of its catalytic domain on amino acids 183 and 185 according to the primary structure. To create a catalytically inactive Map kinase - a kinase dead mutant - the most likely substitution within its active domain would be:
Correct Answer: C
ExplanationProtein phosphorylation is an important regulation of protein activity and function. Upon phosphorylation, a phosphate moiety is covalently attached to a hydroxyl (-OH) group of any of the three amino acids: Tyrosine (Y), Threonine (T) or Serine (S), which are the only 3 amino acids with a side chain containing hydroxyl. Keep in mind that all amino acids between the first and last in a polypeptide or protein, have both their carboxyl and amino ends involved in peptide bonding, so we must assess the side chain.
When a phosphate group is added, a conformational change often occurs within the protein domain. Such conformational change would allow the protein binding its partners and also phosphorylating them to promote a further propagation of protein activation along a signaling pathway. In the mutants suggested in answer choices A, B and D, a partial phosphorylation could still be taking place within the catalytic domain, which could have rendered a partially active Map kinase. However, in the mutant G183A185 (i.e. glycine - G - at position 183 in the primary structure and alanine - A - at position 185) suggested in answer choice C, no phosphorylation is possible within its catalytic domain due to the absence of the hydroxyl functional group, therefore this mutant would be ‘kinase dead’.
Going Deeper: Glycine and alanine are the most commonly used amino acids for substitutions when mutating protein active sites since these amino acids are neutral and relatively small as compared to others.
Background: The new MCAT requires knowledge of the side chains of amino acids, their features, as well as the 3-letter and 1-letter representations. The following image represents the 20 standard amino acids at physiological pH with the 9 essential amino acids identified with a red asterisk.
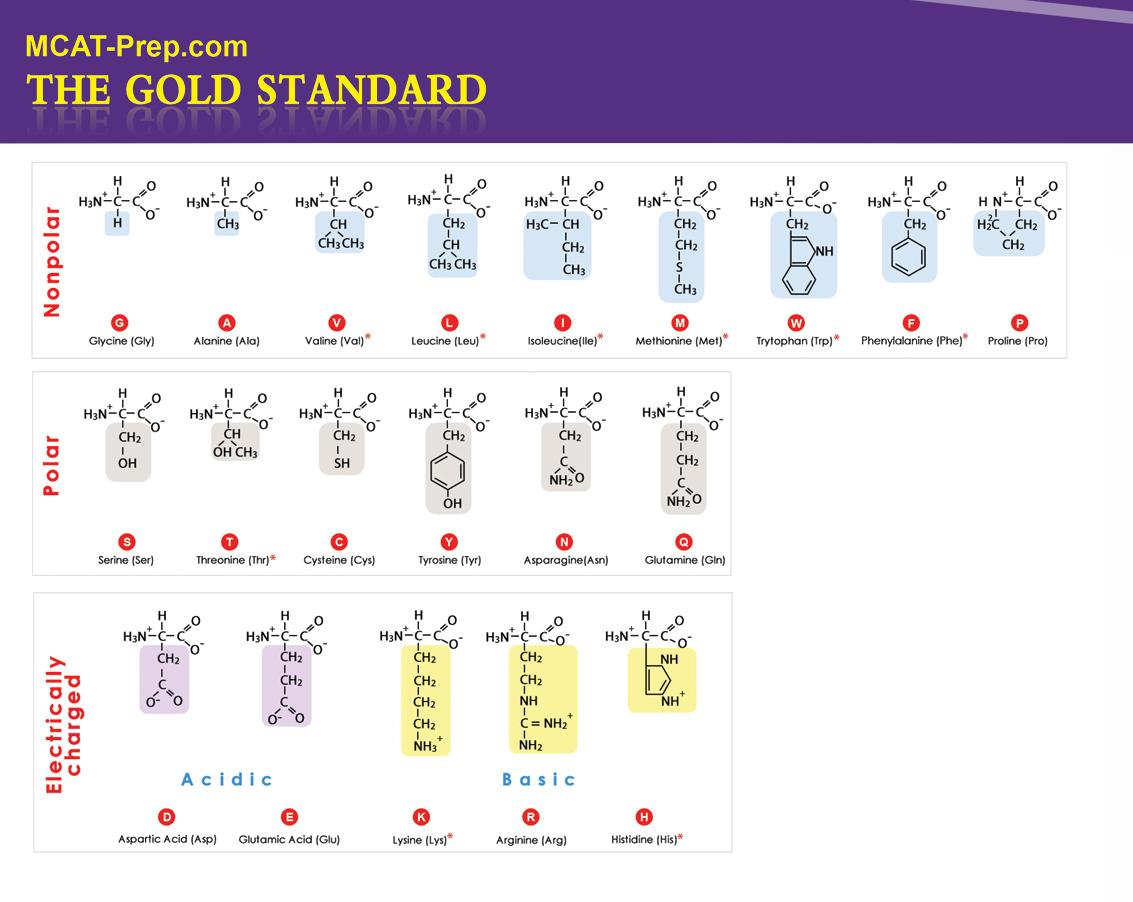

Ala A Alanine Arg R Arginine Asn N Asparagine Asp D Aspartic acid Cys C Cysteine Gln Q Glutamine Glu E Glutamic acid Gly G Glycine His H Histidine Ile I Isoleucine Leu L Leucine Lys K Lysine Met M Methionine Phe F Phenylalanine Pro P Proline Ser S Serine Thr T Threonine Trp W Tryptophan Tyr Y Tyrosine Val V Valine MCAT Biochemistry Practice Question 2
Apoptosis is the process of programmed cell death that can occur in multicellular organisms. The proteins involved in apoptosis are associated with pathways for cell cycle arrest and DNA repair. These processes are mostly regulated through the interplay of various proteins involved in feedback loops including some of the ones shown in Figure 1.
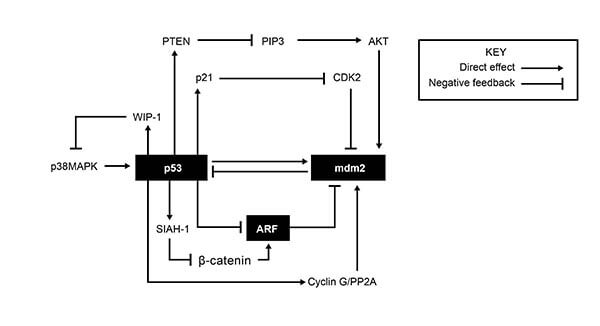
Figure 1: Feedback loops forming a regulatory network affecting apoptosis, cell cycle arrest and DNA repair. (Bioformatics Institute)
According to Figure 1, CDK2 activity would most reasonably increase due to all of the following EXCEPT:
Correct Answer: D
ExplanationNotice the key in the figure which will allow us to follow each arrow that stimulates the next protein and each symbol for negative feedback which means there will be some downregulation (amount/concentration goes down). [Notice a key step in the diagram: p21 inhibits CDK2]
Degradation of p21 implies that the concentration of p21 in its active form goes down. The diagram shows that p21 has a negative influence on CDK2. In other words, when p21 is high, CDK2 goes low. But in our instance, p21 is low (degraded) so this allows CDK2 to rise unchecked.
High cyclin G concentrations: From the bottom of Figure 1, we can see that high cyclin G leads to high mdm2 and low p53 (notice carefully, when we leave mdm2, there is only one place to go in the diagram because all the other symbols are pointing to mdm2 and only one symbol is pointing away). Note that we used the most direct route to get to CDK2 as the question used the words “most reasonably”. Low p53 means low p21 which we established will lead to a rise in CDK2.
A mutation in the gene that produces PTEN: The great majority of mutations will result in an ineffective gene product or none at all. Thus we have a decrease in PTEN which will lead to a rise in PIP3 (if you are unsure, think of what happens if PTEN goes up, then PIP3 must go down because of the negative feedback symbol), rise in AKT, rise in mdm2, decrease in p53 which we already established means an eventual rise in CDK2.
High p53 concentrations: clearly we get the opposite of the above, meaning a decrease in CDK2. High p53 stimulates p21 which has a negative feedback on CDK2.
This is just a taste of what you can expect on the MCAT. Sign up for our FREE MCAT practice test for access to passage-based and discrete practice questions. You'll receive instant scores with detailed explanations, access to helpful science videos for tricky concepts, and gain valuable insights into your strengths and weaknesses. Sign up here: Free MCAT practice test
Gold Standard Science Cheat Sheets
Boost Your Score with These Resources
MCAT Biochemistry Macromolecules: Summary I - Amino Acids and Proteins
The current MCAT regularly has questions which require previous knowledge of the structures, features (including changes in charge with pH), 3- and 1-letter abbreviations of the 20 common protein-generating amino acids, etc. You can consider the following your MCAT Biochemistry cheat sheet (i.e. memory aid for quick reference, but obviously not to be used surreptitiously).
Note: Unless mentioned otherwise, the following images are excerpts from the Gold Standard MCAT Biochemistry ebook.


| Ala | A | Alanine |
| Arg | R | Arginine |
| Asn | N | Asparagine |
| Asp | D | Aspartic acid |
| Cys | C | Cysteine |
| Gln | Q | Glutamine |
| Glu | E | Glutamic acid |
| Gly | G | Glycine |
| His | H | Histidine |
| Ile | I | Isoleucine |
| Leu | L | Leucine |
| Lys | K | Lysine |
| Met | M | Methionine |
| Phe | F | Phenylalanine |
| Pro | P | Proline |
| Ser | S | Serine |
| Thr | T | Threonine |
| Trp | W | Tryptophan |
| Tyr | Y | Tyrosine |
| Val | V | Valine |
Helpful Mnemonics for One-Letter Amino Acid Abbreviations on the MCAT
Alanine, aRginine, asparagiNe, asparDic acid [aspartic acid], Cysteine, Qutamine [glutamine], glutamEc acid [glutamic acid], Glycine, Histidine, Isoleucine, Leucine, Kysine [lysine], Methionine, Fenylalanine [phenylalanine], Proline, Serine, Threonine, tWyptophan [tryptophan], tYrosine, Valine
Nonpolar Amino Acids = G A P V W L I M F or Gap V.W. Lymph
(lymph is important in fatty acid transport . . . think fatty acid tails are nonpolar)
Polar Uncharged Amino Acids = S T Y C N Q or Stick Nick
(stick like a "pole" . . . think polar)
Electric Amino Acids = D E H K R or Dee Hicker
(dee hicker like deelectric . . . think electric)
To be more precise, for the Polar Charged Amino Acids:
Dee Negative, Hicker Positive, D(-) E(-) H(+) K(+) R(+)
General Principles of Proteins
- Primary structure
- Sequence of amino acids encoded by DNA and linked to each other through covalent bonds (including disulfide bonds)
- Secondary structure
- Orderly inter- or intramolecular hydrogen bonding of the protein chain
- Usually organized in a stable ɑ-helix or a β-pleated sheet
- Tertiary structure
- Further folding of the protein molecule onto itself
- 3D shape (spatial organization) of an entire protein molecule
- Quaternary structure
- 2 or more protein chains bonded together by noncovalent bonds
- Isoelectric point (pI)
- pH at which a given amino acid will be neutral (have no net charge)
- Average of the 2 pKa values of an amino acid (depending on the dissociated group)
- Both amino acids and proteins are least soluble at their isoelectric points
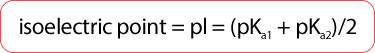


Levels of Protein Organization
[Wikipedia, Creative Commons]
Macromolecules: Summary II - Carbohydrates
- Compounds consisting of aldehydes and ketones with general formula Cm(H2O)n
- Monosaccharides = simplest carbohydrate unit
- Can be named according to number of carbons it contains
- Common names: triose (3 carbons), tetrose (4 carbons), pentose (5 carbons), hexose (6 carbons)
- Note: '-ose' suffix denotes sugar
- Aldose = aldehyde sugar
- Ketose = ketone sugar
- Disaccharides = two sugars bound together
- Polysaccharides = long sugar chains for glucose storage (i.e. glycogen in animals, starch in plants)
- Absolute configuration
- Describes spatial arrangement of atoms or groups around a chiral molecule
- Assign priority to substituents of the chiral carbon
- The higher the atomic number, the higher the priority
- Orient the molecule with the lowest priority substituent in the back
- If the priorities increase in a clockwise direction = R
- If the priorities decrease in a clockwise direction = S
- Cyclic structure
- Monosaccharides can undergo intramolecular reactions to form ring structures
- Cyclic sugars are stable in solution and can form two types of rings:
- Six-membered rings (6 carbons in ring) = pyranose
- Five-membered rings (5 carbons in ring) = furanose
- Glycosidic linkages
- Covalent bonds between monosaccharides and alcohols
- When alcohol is another monosaccharide, produces a disaccharide
- i.e. sucrose, lactose, maltose, cellobiose
- Linkage between C1 on the first sugar and C2 on the second sugar = 1,2 linkage
- Linkage between C1 on the first sugar and C4 on the second sugar = 1,4 linkage
- Linkage between C1 on the first sugar and C6 on the second sugar = 1,6 linkage
- May also be classified as alpha (α) or beta (β):
- Hydroxyl group on C1 oriented up = beta (βirds fly in the sky)
- Hydroxyl group on C1 oriented down = alpha (α - fish - swim in the sea)
- Epimers vs. Anomers
- Epimers - differ in absolute configuration at only one carbon (R or S)
- Anomers - differ in configuration at the hemiacetal/acetal (= anomeric) carbon
- Note: Conversion between anomers = mutarotation

D, R-glyceraldehyde


Macromolecules: Summary III - Fatty Acids
- Amphipathic molecules possessing a polar carboxylate-head group and nonpolar hydrocarbon tail
- Saturated fatty acids
- Maximum hydrogens - stack together
- No double bonds
- Solids at room temperature (= fats; example: butter)
- Higher melting point, boiling point, and Van der Waals
- Unsaturated fatty acids
- 1 or more double bonds - bent or kinked structure
- Liquids at room temperature (= oils; example: olive oil)
- Exist in body in 3 common forms:
- Triglycerides (also known as triacylglycerols)
- 3 fatty acids esterified to glycerol
- Nonpolar tails - used for energy storage
- Digested by lipase
- With low food intake, broken down by liver to form ketone bodies
- Phospholipids
- Fatty acids bound to glycerol, phosphate, and other groups
- Amphipathic (hydrophilic + hydrophobic components) - used for cell membranes
- Cholesterol
- Fatty acids in ring form [isoprene units derived from acetyl-CoA cyclize to form a four-ringed structure]
- Cholesterol is amphipathic. The ring is nonpolar and the OH-head group is polar.
- Used for steroid hormones, bile, membrane fluidity
- Triglycerides (also known as triacylglycerols)
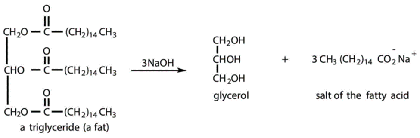

The Three Ketone Bodies

Basic Ring Structure of Steroids
Macromolecules: Summary IV - Nucleic Acids
- Nucleotides (also known as nucleoside phosphates)
- are composed of a 5-carbon sugar, a nitrogenous base, and an inorganic phosphate (a glycosidic bond links the nitrogenous base to the sugar)
- are attached in sequence via phosphodiester bonds to form nucleic acids
- form the sugar phosphate (negatively-charged) backbone of nucleic acids
- Nucleosides
- are composed of a nitrogenous base and a 5-carbon sugar
- are attached to an inorganic phosphate to form nucleotides
- Nitrogenous Bases
- Purines
- 2 rings - adenine (A) and guanine (G) - mnemonic: Pure As Gold
- Pyrimidines
- 1 ring - cytosine (C), uracil (U), thymine (T) - mnemonic: CUT the Py
- Purines
Note the direction of hydrogen bonding in the diagram below:


- Watson–Crick model of DNA
- Double helical structure composed of 2 complementary and anti-parallel DNA strands held together by hydrogen bonds between nitrogenous base pairs
- A binds T with 2 hydrogen bonds; C binds G with 3 hydrogen bonds
- Mnemonic: All Tigers and Cats Growl
- Nitrogenous bases project to center of double helix to hydrogen bond with each other (think of double helix as a winding staircase - each stair represents a nitrogenous base pair keeping the helix shape intact)
- Double helical structure composed of 2 complementary and anti-parallel DNA strands held together by hydrogen bonds between nitrogenous base pairs
| Deoxyribonucleic Acid (DNA) | Ribonucleic Acid (RNA) |
| Contains the genetic information of the cell - transfers that genetic information | Helps DNA transfer genetic information for creation of proteins |
| Made from deoxyribose | Made from ribose |
| Double-stranded | Mostly single-stranded |
| Adenine binds Thymine Cytosine binds Guanine |
Adenine binds Uracil Cytosine binds Guanine |
| Found in nucleus and mitochondria | Found in nucleus, cytoplasm, and ribosomes |
| Replicated by DNA polymerases | Transcribed from DNA |
- DNA Denaturation
- Separation of DNA strands due to high temperatures, chemicals (i.e. urea), or UV radiation
- Occurs when hydrogen bonds between base pairs are broken
- Separated strands are still complementary
- DNA Reannealing (also known as hybridization)
- Single, complementary strands stick tightly and reform double-stranded DNA
- Single strands of DNA can only hybridize with each other if they are complementary

MCAT Biochemistry Kinetics: Summary I
- Kinetics tell how fast a reaction will occur
- Catalysts
- speed up reactions by speeding up the rate-determining step or providing a different route to the products; lower the energy of activation
- remain unchanged at the end of the reaction
- are not included in the overall reaction equation

- Michaelis-Menten
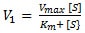
- V0 = initial velocity (reaction rate)
- Vmax = maximum reaction rate
- [S] = concentration of substrate
- Km = amount of substrate needed for enzyme to obtain half of its maximum rate of reaction (Michaelis-Menten constant)
- Low Km = high affinity for substrate
- High Km = low affinity for substrate
- Ka = association constant of the enzyme-substrate complex
- Low Ka = low affinity for substrate (enzyme-substrate complex is less stable)
- High Ka = high affinity for substrate (enzyme-substrate complex is more stable)
- Kd = dissociation constant of the enzyme-substrate complex
- Low Kd = high affinity for substrate (enzyme-substrate complex is more stable)
- High Kd = low affinity for substrate (enzyme-substrate complex is less stable)
- kcat = number of substrate molecules each enzyme converts to product per unit time (turnover number)
- Low kcat = enzyme-substrate complex converts less of substrate it binds into product
- High kcat = enzyme-substrate complex converts more of substrate it binds into product
- Feedback Regulation
- Positive Feedback - promotes or enhances reactions
- Negative Feedback - inhibits or reduces reactions
- Cooperativity (also known as positive or negative cooperative binding)
- Occurs when binding of Substrate B to Substrate A increases (positive) or decreases (negative) activity of Substrate A
- Michaelis-Menten kinetics do not apply to enzymes engaging in cooperative binding
MCAT Biochemistry Kinetics: Summary II - Enzymes
- Competitive Inhibitors
- Compete with the substrate in binding to the enzyme's active site
- Are structurally similar to the substrate
- Bind to free enzyme (= E) only forming an unreactive enzyme-inhibitor complex (= EI)
- Cannot bind to the enzyme at the same time as the substrate
- Can only bind if the substrate has not bound
- Are reversible with increasing [S]
- Increase Km and have no effect on Vmax
- Uncompetitive Inhibitors
- Bind to the enzyme-substrate complex (= ES) only forming an unreactive enzyme-inhibitor-substrate complex (= EIS)
- Can only bind if the substrate has bound
- Are reversible with decreasing [S]
- Decrease Km and decrease Vmax
- Mixed Inhibitors
- Bind to the enzyme at a site other than the active site
- Bind to free enzyme (= E) or the enzyme-substrate complex (= ES) forming an unreactive enzyme-inhibitor complex (= EI) or enzyme-inhibitor-substrate complex (= EIS), respectively
- Can bind whether or not the substrate has bound
- Produce a conformational change in the enzyme - binding of the inhibitor reduces the substrate's affinity for the active site
- Are reduced, but not reversible with increasing [S]
- Increase or decrease Km and decrease Vmax
- Noncompetitive Inhibitors
- Do not compete with the substrate in binding to the enzyme's active site
- Bind to the enzyme at a site other than the active site
- Bind to free enzyme (= E) or the enzyme-substrate complex (= ES) forming an unreactive enzyme-inhibitor complex (= EI) or enzyme-inhibitor-substrate complex (= EIS), respectively
- Can bind whether or not the substrate has bound
- Are not reduced or reversible with increasing [S] - must remove inhibitor
- Decrease Vmax and have no effect on Km
Note: Noncompetitive inhibition is sometimes considered a special case of mixed inhibition. Noncompetitive inhibitors have the same affinity for free enzyme (= E) and the enzyme-substrate complex (= ES) whereas mixed inhibitors tend to have a higher affinity for either free enzyme (= E) or the enzyme-substrate complex (= ES).


[Wikipedia, Creative Commons]
| Inhibitor | Binds | Reversible? | Effect |
| Competitive | Free enzyme (E) | Yes with ↑[S] | Increases Km |
| Uncompetitive | Enzyme-substrate complex (ES) | Yes with ↓[S] | Decreases Km and Vmax |
| Mixed | Free enzyme (E) or enzyme-substrate complex (ES) | Reduced with ↑[S] | Increases or decreases Km and decreases Vmax |
| Noncompetitive | Free enzyme (E) or enzyme-substrate complex (ES) | Yes with removal of inhibitor | Decreases Vmax |
KoMpetitive INhibition = KM INcrease (Vmax is unchanged)
NOn-KoMpetitive INhibition = NO KM INcrease (but Vmax is decrease)
Uncompetitive Inhibition = BOTH Km and Vmax decrease
Mnemonic for Remembering the Effects of Enzyme Inhibitors
- Ki = binding affinity of the inhibitor
- Ki > 1 - inhibitor has higher affinity for enzyme than substrate
- Ki < 1 - inhibitor has lower affinity for enzyme than substrate
- IC50 = half-maximal inhibitory concentration
- Tells how much of a drug is needed to inhibit a biological process by 50%
- Allosteric enzymes have 2 configurations:
- Active state - catalyzes reactions
- Inactive state - cannot catalyze reactions
- Zymogens are inactive enzymes that must be cleaved to become active


Thermodynamics: Summary
- Thermodynamics tell if a reaction will occur
- Gibbs Free Energy (ΔG) - tells nothing about rate of reaction, only its spontaneity
- ΔG = ΔH - TΔS
- Negative ΔG = exergonic = spontaneous
- ΔG = 0 = equilibrium
- Positive ΔG = endergonic = nonspontaneous
- Equilibrium constant (Keq)
- Keq = products/reactants
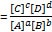 for a reaction A + B → C + D
for a reaction A + B → C + D
- Note: a, b, c, and d are stoichiometries of the substrates and products, respectively
- Keq > 1 - reaction favors products
- Keq < 1 - reaction favors reactants
- Keq = products/reactants
- Gibbs Free Energy at standard state (ΔGo) = 298K (25oC), 1 atm, pure solids/liquids, 1M solutions
- ΔGo < 0 and Keq > 1 . . . products are favored
- ΔGo = 0 and Keq = 1 . . . products are approx. equal to reactants
- ΔGo > 0 and Keq < 1 . . . reactants are favored
- Le Chatelier's Principle
- Reaction in equilibrium will change direction according to applied stress
- Stress could be a change in concentration, pressure, temperature, or volume
- Enthalpy
- Measure of heat energy absorbed or released when bonds are broken or formed, respectively, during a reaction run at constant pressure
- Negative ΔH - exothermic - bond(s) formed - energy released
- Positive ΔH - endothermic - bond(s) broken - energy absorbed
| ΔH | ΔS | ...ΔG | Reaction Spontaneity |
| - | + | - | Spontaneous at all temperatures |
| - | - | - or + | Spontaneous at low temperatures where ΔH outweighs TΔS Nonspontaneous at high temperatures where TΔS outweighs ΔH |
| + | - | + | Nonspontaneous at all temperatures |
| + | + | - or + | Spontaneous at high temperatures where TΔS outweighs ΔH Nonspontaneous at low temperatures where ΔH outweighs TΔS |

Plasma Membrane: Summary I
- A phospholipid bilayer that encloses cells
- Outer region - hydrophilic, polar, phosphoric acids (= heads)
- Inner region - hydrophobic, nonpolar, fatty acids (= tails)
- Cholesterol found within membrane for structural support and as a fluidity buffer
- Transmembrane proteins assist in solute transport across membranes
| Process (solute type) | Concentration Gradient | Requires Protein? | Requires Energy? |
| Diffusion (small nonpolar) | High to Low | No | No – passive |
| Osmosis (water only) | High to Low | No | No – passive |
| Facilitated Transport (large nonpolar) | High to Low | Yes | No – passive |
| Active Transport (polar/ions) | Low to High | Yes | Yes – active (ATP) |
Note: Difference in polarity between (+) outside and (-) inside along with solute transport creates a membrane potential across the cell membrane.
Osmotic pressure = pressure required to resist movement of water through a semipermeable membrane due to concentration gradient
Plasma Membrane: Summary II
- Exocytosis
- Vesicle inside cell fuses with cell membrane and releases contents to outside
- Endocytosis
- Cell membrane bends inward to form vesicle around particles or liquids
- If large particles = phagocytosis
- If small particles or liquids = pinocytosis
- Cell membrane bends inward to form vesicle around particles or liquids
- Note: Contents never cross cell membrane in either process
- Biosignaling
- Ion Channels - allow for passive or active transport of ions; two types:
- Voltage-gated - transmembrane proteins activated by changes in membrane potential
- Ligand-gated - transmembrane proteins activated by binding of a specific ligand
- Ion Channels - allow for passive or active transport of ions; two types:
- Receptor enzymes (also known as enzyme-linked receptors)
- Transmembrane proteins
- Binding of ligand on extracellular side produces activity on the intracellular side
- G protein-coupled receptors
- Integral membrane proteins
- Sensing of molecules outside cell activates response inside cell

Amphipathic molecules arranged in micelles, a liposome and a bilipid layer [= plasma membrane]
Biotechnology: Summary I
- DNA recombination
- Involves combining DNA segments or genes from different sources
- Foreign DNA can come from another DNA molecule, a chromosome, or a complete organism
- Uses restriction enzymes from bacteria that cut pieces of DNA at specific recognition sites (= nucleotide sequences) along the DNA strand
- When a double-stranded DNA segment is cut, sticky ends are produced
- Sticky ends = unpaired, single-strand extensions of DNA that are ready to bind with complementary codons (= sequence of 3 adjacent nucleotides)
- The cut pieces of DNA = restriction fragments
- Restriction fragments are then inserted into plasmids
- Plasmids (also known as replicons)
- Circular pieces of DNA that replicate independently of chromosomal DNA
- Cut with the same restriction enzymes as the restriction fragment to generate the same sticky ends
- Plasmids (also known as replicons)
- Plasmids are introduced into the bacteria via transformation
- Restriction fragments and linearized plasmids are joined together by DNA ligase to form a recombinant plasmid
- Special application
- DNA recombination makes it possible for bacteria to make many copies of recombinant plasmids and produce proteins that are not native to its species (e.g. bacteria with recombinant DNA producing insulin to treat diabetes)

- More on restriction enzymes (also known as restriction endonucleases - endo = internal; nuclease = cut DNA)
- Naturally found in bacteria and archaea
- Serve as the organism's own natural defense mechanism
- Enzymes cleave foreign DNA at specific recognition sites when it enters a bacterium
- The organism's own DNA is protected by selective methylation at these sites via methylase enzymes
- Come in 3 types - classified by their structure and how they cleave their DNA substrate
- Type I, II, and III – Type II is the most common
- Type II restriction enzymes
- Cleave within or near specific recognition sites (short, specific, palindromic DNA sequences where cleavage occurs under the right conditions)
- Note: palindromic sequences read the same forward or backward (e.g. RACE CAR)
- Gene cloning
- Involves the use of a cloning vector (e.g. plasmid, bacteriophage, virus, etc.)
- All cloning vectors must contain at least one unique cloning site
- That cloning site must be cleavable by a restriction enzyme as well as a place where a gene of interest can be inserted
- Multiple Cloning Site
- An artificially-engineered region of several cloning sites lumped together
- Found in many commercial and artificially modified cloning vectors
- Allows the researcher to pick restriction sites that:
- are unique to the vector
- ensure that translocation occurs in-frame
- Origin of replication (ori) = allows cloning vectors to self-replicate
- Special applications
- Cloning vectors are often modified to carry genes that provide resistance to antibiotics, like ampicillin or kanamycin, which allows vectors to be used for selecting certain bacteria
- Bacteriophage vectors have high transfection (= introduction of nucleic acids into cells) rates and are easy to screen which makes them useful for large-scale applications (e.g. genomic DNA screening)
- DNA libraries
- are collections of recombinant DNA
- can consist of (1) genomic DNA or (2) cDNA
- Genomic DNA library
- Contains recombinant DNA that represents the entire genome of an organism
- How it is generated:
- Partially digest genomic DNA with restriction enzymes to ensure representation of all genes
- Digests bacteriophage vector with the same enzyme
- Ligate digested genomic DNA with vector - pack both into bacteriophages
- Infect densely populated E.coli with bacteriophages
- Isolate cleared areas of lysed cells (= plaques)
- cDNA (= complementary DNA) library
- Contains double-stranded (= dsDNA) that is reverse transcribed from messenger RNA (= mRNA)
- How it is generated (in eukaryotes):
- Purify mRNA by applying it to an oligodeoxythymidylate (oligo dT) cellulose filter which binds the mRNA's poly A-tail
- Synthesize cDNA using oligo dT as a primer
- Cut mRNA with RNA-nicking RNase H and DNA polymerase I uses the fragments as primers to synthesize cDNA
- Digest resulting double-stranded cDNA and insert it into vectors
- Create phage library as discussed for genomic DNA library
- Unlike genomic DNA libraries, an advantage of cDNA libraries is that they do not include introns or flanking sequences
- Locating a gene of interest
- Use a hybridization probe
- A molecule that recognizes a specific DNA sequence or protein product labeled with a compound that allows for easy detection (e.g. radioactive isotope, reactive enzyme that produces color, or fluorescent dye)
- Hybridize the probe and the DNA library
- The sensitivity or stringency of the hybridization reaction can be increased or decreased by changing experimental variables (e.g. salt concentration, temperature)
- Isolate detected positive hits from phage plates
- Identify and isolate the gene of interest
- Amplify the gene using polymerase chain reaction (PCR)
- Use a hybridization probe
- Polymerase Chain Reaction (PCR)
- Allows for the rapid amplification of any DNA fragment without purification
- DNA primers flank the specific sequence to be amplified
- Primers are then extended to the end of the DNA molecule with the use of a heat-resistant DNA polymerase
- Newly synthesized DNA strand is then used as the template to undergo another round of replication
- Technique:
- Melt target DNA into two single strands by heating the reaction mixture to approximately 94 °C
- Cool the mixture rapidly to anneal the primers to their specific locations
- Note: anneal = the sticking together of complementary single strands via the formation of hydrogen bonding between base pairs
- Raise the temperature to 72°C to allow DNA polymerase to add nucleotides until the entire complementary strand of the template is completed
- Repeat the cycle to make more copies
- Special applications:
- Sex determination
- Requires amplification of intron 1 of amelogenin gene
- Amelogenin gene, found on X-Y homologous chromosomes, contains a 184 base pair deletion on the Y homologue
- That deletion allows females to be distinguished from males in that males will have two different sizes of amplified DNA whereas females will have one unique size
- Introduce specific mutations into genes for research purposes such as:
- Deleting entire fragments
- Changing specific amino acids
- Adding peptide tags (= peptide sequences tagged onto a protein)
- Sex determination

- DNA Sequencing
- Method by which to ensure that the gene insert of interest does not contain any mistakes
- Sanger method of DNA sequencing
- Most commonly used sequencing technique
- Makes four preparations of DNA
- Each preparation is incubated with a labeled sequencing primer, DNA polymerase, all 4 DNA triphosphates, and 1 of the nucleotides in its dideoxy form (e.g. ddATP, ddCTP, ddGTP, or ddTTP)
- Technique:
- New strands of DNA are synthesized using PCR beginning at the primer and continuing on until a dideoxynucleotide (ddNTP) is inserted at random
- This results in a mixture of DNA fragments with varying lengths that must be separated by gel electrophoresis and read one nucleotide at a time

MCAT Biochemistry Biotechnology: Summary II
- Gel electrophoresis
- Method of separating DNA fragments of differing lengths based on their size
- Technique:
- DNA fragments are passed through an electrically-charged agarose gel
- Since DNA is negatively charged, it moves towards the anode (= positive electrode)
- Shorter fragments move faster than the longer fragments and can be visualized as a banding pattern using autoradiography techniques
- Southern blotting
- Method of transferring DNA fragments from the electrophoresis agarose gel onto filter paper where they are identified with probes
- Technique:
- DNA is digested in a mixture with restriction endonucleases to cut out specific pieces of DNA
- The DNA fragments are then subjected to gel electrophoresis
- Separated fragments are bathed in an alkaline solution where they immediately begin to denature
- Fragments are then blotted onto nitrocellulose paper and incubated with a specific probe whose location can be visualized with autoradiography
- Northern blotting
- Method of detecting specific sequences of RNA by hybridization with cDNA
- Western blotting
- Method used to identify specific amino acid sequences in proteins
- Proteins are separated by gel electrophoresis on an SDS-PAGE gel and detected using labeled antibodies specific to the protein or part of the protein


Mnemonic for Remembering the Blotting Techniques
- DNA microarrays (= DNA chip or biochip or "laboratory-on-a-chip")
- Determine which genes are active or inactive in different cell types
- Evolved from Southern blotting
- Can be used to genotype multiple regions of a genome
- Created by robotic machines that arrange small amounts of hundreds of thousands of gene sequences on a single microscope slide
- Sequences can be a short section of a gene or other DNA element that is used to hybridize a cDNA or cRNA (also called antisense RNA) sample
- Hybridization is usually observed and quantified by the detection of a fluorescent tag
- Protein Quantification
- Quantitative analysis techniques allow researchers to determine basic information about the expressed product (e.g. concentration, amount, size)
- SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis)
- Method of separating proteins according to their electrophoretic mobility
- SDS is an anionic (= negatively charged) detergent with the following effects: (1) denaturing proteins and (2) giving an additional negative charge to the now-linearized proteins
- In most proteins, the binding of SDS to the polypeptide chain gives an even distribution of charge per unit mass, thus fractionation will approximate size during electrophoresis
- Column Chromatography
- Method of purifying proteins according to their size, charge, hydrophobicity, and more
- Affinity Chromatography
- Allows the rapid purification of biological molecules based on certain affinities
- Examples: affinities between antigen and antibody, enzyme and substrate, or genetically engineered protein tags and specific matrices
- Ion Chromatography
- Allows the purification of biological molecules based on overall charge:
- Cation Chromatography - selectively binds positively-charged molecules
- Anion Chromatography - selectively binds negatively-charged molecules
- Proteins are removed by washing (= eluting) using increasing concentrations of salt
- Allows the purification of biological molecules based on overall charge:
- Size-Exclusion Chromatography (SEC) (also known as gel filtration)
- Allows the purification of biological molecules based on size
- A mixed protein solution is applied to the top of a long column packed with porous polymer beads
- As the solution filters through the beads, smaller molecules are slowed down by the pores in the beads while larger molecules are allowed to flowed through at a faster rate - thus, larger molecules are eluted first

MCAT Biochemistry Practice Questions
These MCAT Biochemistry practice questions are taken from our Gold Standard MCAT Question Bank that contains over 4000 practice questions and growing. These questions were chosen to give you a sense as to the reasoning and knowledge required in Biochemistry for the current MCAT exam. It would be better to complete some part of your review before attempting these practice questions. Good luck!
MCAT Biochemistry Practice Question 1
In order to become active, a Map kinase has to undergo phosphorylation of its catalytic domain on amino acids 183 and 185 according to the primary structure. To create a catalytically inactive Map kinase - a kinase dead mutant - the most likely substitution within its active domain would be:
Explanation
Protein phosphorylation is an important regulation of protein activity and function. Upon phosphorylation, a phosphate moiety is covalently attached to a hydroxyl (-OH) group of any of the three amino acids: Tyrosine (Y), Threonine (T) or Serine (S), which are the only 3 amino acids with a side chain containing hydroxyl. Keep in mind that all amino acids between the first and last in a polypeptide or protein, have both their carboxyl and amino ends involved in peptide bonding, so we must assess the side chain.
When a phosphate group is added, a conformational change often occurs within the protein domain. Such conformational change would allow the protein binding its partners and also phosphorylating them to promote a further propagation of protein activation along a signaling pathway. In the mutants suggested in answer choices A, B and D, a partial phosphorylation could still be taking place within the catalytic domain, which could have rendered a partially active Map kinase. However, in the mutant G183A185 (i.e. glycine - G - at position 183 in the primary structure and alanine - A - at position 185) suggested in answer choice C, no phosphorylation is possible within its catalytic domain due to the absence of the hydroxyl functional group, therefore this mutant would be ‘kinase dead’.
Going Deeper: Glycine and alanine are the most commonly used amino acids for substitutions when mutating protein active sites since these amino acids are neutral and relatively small as compared to others.
Background: The new MCAT requires knowledge of the side chains of amino acids, their features, as well as the 3-letter and 1-letter representations. The following image represents the 20 standard amino acids at physiological pH with the 9 essential amino acids identified with a red asterisk.

| Ala | A | Alanine |
| Arg | R | Arginine |
| Asn | N | Asparagine |
| Asp | D | Aspartic acid |
| Cys | C | Cysteine |
| Gln | Q | Glutamine |
| Glu | E | Glutamic acid |
| Gly | G | Glycine |
| His | H | Histidine |
| Ile | I | Isoleucine |
| Leu | L | Leucine |
| Lys | K | Lysine |
| Met | M | Methionine |
| Phe | F | Phenylalanine |
| Pro | P | Proline |
| Ser | S | Serine |
| Thr | T | Threonine |
| Trp | W | Tryptophan |
| Tyr | Y | Tyrosine |
| Val | V | Valine |
MCAT Biochemistry Practice Question 2
Apoptosis is the process of programmed cell death that can occur in multicellular organisms. The proteins involved in apoptosis are associated with pathways for cell cycle arrest and DNA repair. These processes are mostly regulated through the interplay of various proteins involved in feedback loops including some of the ones shown in Figure 1.

Figure 1: Feedback loops forming a regulatory network affecting apoptosis, cell cycle arrest and DNA repair. (Bioformatics Institute)
According to Figure 1, CDK2 activity would most reasonably increase due to all of the following EXCEPT:
Explanation
Notice the key in the figure which will
allow us to follow each arrow that stimulates the next protein and each symbol for negative feedback
which means there will be some downregulation (amount/concentration goes down). [Notice a key step in
the diagram: p21 inhibits CDK2]
Degradation of p21 implies that the concentration of p21 in its active form goes down. The diagram
shows that p21 has a negative influence on CDK2. In other words, when p21 is high, CDK2 goes low. But in
our instance, p21 is low (degraded) so this allows CDK2 to rise unchecked.
High cyclin G concentrations: From the bottom of Figure 1, we can see that high cyclin G leads to high
mdm2 and low p53 (notice carefully, when we leave mdm2, there is only one place to go in the diagram because
all the other symbols are pointing to mdm2 and only one symbol is pointing away). Note that we used the most
direct route to get to CDK2 as the question used the words “most reasonably”. Low p53 means low p21 which we established
will lead to a rise in CDK2.
A mutation in the gene that produces PTEN: The great majority of mutations will result in an ineffective gene product
or none at all. Thus we have a decrease in PTEN which will lead to a rise in PIP3 (if you are unsure, think of what happens
if PTEN goes up, then PIP3 must go down because of the negative feedback symbol), rise in AKT, rise in mdm2, decrease in p53
which we already established means an eventual rise in CDK2.
High p53 concentrations: clearly we get the opposite of the above, meaning a decrease in CDK2. High p53 stimulates p21 which
has a negative feedback on CDK2.